A Chemical Fingerprint of Everything You’ve Taken and Forgotten
Which medications have you taken, and might you have forgotten some of them? What other substances could be present in the body that patients are unaware of?
That question still underpins a large part of clinical research today. Information about drug exposure is most often obtained from medical records or patient self-reports. However, such sources are frequently incomplete: people forget what they have taken, use over-the-counter medications, return to old prescriptions, or are exposed to compounds through food and the environment. As a result, the true picture of what the body has been exposed to is often incomplete.
To obtain a more direct and comprehensive view, researchers developed the GNPS (Global Natural Product Social Molecular Networking) Drug Library, an extensive reference database based on mass spectrometry (MS/MS), which enables the direct detection of drugs and their derivatives in biological samples. They analyzed thousands of public metabolomics datasets and identified drug-related compounds and their breakdown products.
The results are presented in the study “A resource to empirically establish drug exposure records directly from untargeted metabolomics data.” We spoke with senior co-author Pieter Dorrestein, Ph.D., professor at the UC San Diego Skaggs School of Pharmacy and Pharmaceutical Sciences and professor of pharmacology and pediatrics at the UC San Diego School of Medicine.
The tool can be applied across multiple scales, from individual patients to population-level trends in wastewater
A key strength of this approach lies in data integration, which enables drugs and related compounds to be identified and linked to information such as their chemical identity and function. The application of this library, however, extends beyond clinical samples. As the authors note, „GNPS Drug Library can be applied for wastewater-based epidemiology to monitor population-level health trends. In wastewater samples collected from March to June 2020, we observed seasonality of drug usage: the abundances of a cough suppressant (dextromethorphan) and antibiotics decreased over the months, while those of antihypertensives, antidepressants, HIV medications, and antiepileptics remained stable. GNPS Drug Library can also be applied in the context of food monitoring. Using untargeted metabolomics files from ~3500 foods/beverages collected in the Global FoodOmics Project82, we observed antibiotics (e.g., ampicillin, tetracycline) in fish, beef, and turkey, as well as antiparasitic drugs (e.g., spinosad, thiabendazole) that are also used as fungicides or insecticides in fruit and vegetables. The mass spectrometry community will play a key role in the evolution of this resource through the continued deposition of reference libraries and expansion of the public metabolomics datasets for analog searches.“
These examples illustrate how the same tool can be applied across multiple scales from individual patients to population-level trends in wastewater, and even to food and the broader environment. At each of these levels, the GNPS Drug Library enables chemical traces of drugs to be translated into biological and societal contexts of exposure. However, as the study emphasizes, all identifications remain based on spectral matching and require additional validation when used for clinical interpretation.
Data science can help to track exposure to antimicrobials, but determining if this results in resistance would require additional follow-up experiments
What was the most difficult part of conducting this research?
Pieter Dorrestein: The most difficult part was building the reference libraries (fingerprints) and the curation of the metadata connected to the MS/MS spectra. This is because there are 1.7 million reference spectra in the public domain, many of which have many names. Penicillin, for example, has several hundred names, so it is not always easy to recognize. Take (2S,5R,6R)-3,3-dimethyl-7-oxo-6-[(phenylacetyl)amino]-4-thia-1-azabicyclo[3.2.0]heptane-2-carboxylic acid; the systematic, internationally recognized, standardized IUPAC name for penicillin is (2S,5R,6R)-3,3-dimethyl-7-oxo-6-[(phenylacetyl)amino]-4-thia-1-azabicyclo[3.2.0]heptane-2-carboxylic acid. We then had to do a literature search, search drug databases, and web searches to connect the molecule names or structural names to find indications and mechanisms of action.
A lot of publicly available data is faulty, so it is important to have the data curated by experts in the field of pharmacy. Kine Eide Kvitne, one of the first authors, and co-senior author Shirley Tsunoda are trained pharmacists who were essential for cleaning up the metadata. Once curated, we could now go into public metabolomics data to discover fingerprints of metabolized versions in public data.
Your study comes at a very important time, so I would like to ask: how significant are these findings in the context of antibiotic resistance?
Pieter Dorrestein: For me, the application of this library will be useful for understanding drug exposure, including antibiotics. We see quite a lot of antimicrobials in food – including dairy, meats, fruits, and vegetables. For example, the antifungal compound natamycin, used to treat eye infections, is often added to dairy products, including organic dairy products, and prevents fungal growth, in particular at the surface (and is used in Europe and the U.S.). So data science can help to track exposure to antimicrobials, but determining if this results in resistance would require additional follow-up experiments.
“Testing more than 3,000 food products, the team found antibiotics in meat products and a pesticide in vegetables that are also used in humans. They believe the library will also be useful for uncovering hidden environmental drug exposures, such as those in reclaimed water and snow.” How can these findings be best applied in practice?
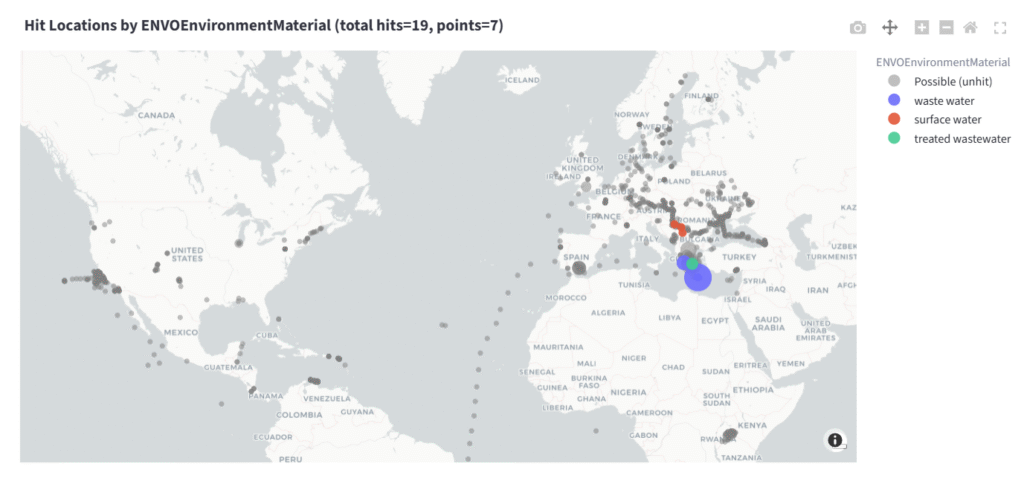
Pieter Dorrestein: In principle, yes. One simply needs to collect untargeted metabolomics data that generates MS/MS (fingerprints) and then just match against this library. It will be important, however, to make sure you then follow up and match the data against a standard to confirm and quantify to ensure that the annotations are correct. Here is an example of the fingerprint of acetaminophen and the environmental or wastewater samples in which it is found.
As they explain, the GNPS Drug Library lays the foundation for future studies that connect drug exposure, microbial breakdown products, and patient outcomes, such as why not all patients respond to treatment in the same way, or how drugs are metabolized and broken down. According to co-first author Nina Zhao, Ph.D., a postdoctoral scientist in the laboratory of Professor Pieter Dorrestein, this could help optimize drug-based therapies. If we do not know what we are exposed to, we cannot adequately respond to the potential threats those substances pose. The GNPS Drug Library provides exactly this necessary visibility, transforming complex chemical signals into clear, usable information.
Image: Pieter Dorrestein, Ph.D. Photo by Kyle Dykes/UC San Diego Health Sciences.
The results are presented in the study “A resource to empirically establish drug exposure records directly from untargeted metabolomics data”, authored by: Haoqi Nina Zhao, Kine Eide Kvitne, Corinna Brungs, Siddharth Mohan, Vincent Charron-Lamoureux, Wout Bittremieux, Runbang Tang, Robin Schmid, Santosh Lamichhane, Shipei Xing, Yasin El Abiead, Mohammadsobhan S. Andalibi, Helena Mannochio-Russo, Madison Ambre, Nicole E. Avalon, MacKenzie Bryant, Lindsey A. Burnett, Andrés Mauricio Caraballo-Rodríguez, Martin Casas Maya, Loryn Chin, Lluís Corominas, Ronald J. Ellis, Donald Franklin, Sagan Girod, Paulo Wender P. Gomes, Lauren Hansen, Robert K. Heaton, Jennifer E. Iudicello, Alan K. Jarmusch, Lora Khatib, Scott Letendre, Sarolt Magyari, Daniel McDonald, Ipsita Mohanty, Andrés Cumsille, David J. Moore, Prajit Rajkumar, Dylan H. Ross, Harshada Sapre, Mohammad Reza Zare Shahneh, Ruben Gil-Solsona, Sydney P. Thomas, Caitlin Tribelhorn, Helena M. Tubb, Corinn Walker, Crystal X. Wang, Jasmine Zemlin, Simone Zuffa, David S. Wishart, Pablo Gago-Ferrero, Rima Kaddurah-Daouk, Mingxun Wang, Manuela Raffatellu, Karsten Zengler, Tomáš Pluskal, Libin Xu, Rob Knight, Shirley M. Tsunoda, and Pieter C. Dorrestein.
The study was funded, in part, by the National Institutes of Health (NIH) (grants 1U19AG063744, 3U19AG063744, P50HD106463, R01DK136117, U24DK133658, P30MH062512) and the Gordon and Betty Moore Foundation (grant GBMF12120).


